image source wikipedia
DBT BET JRF Exam, 2017
Model Question Paper with Answer Key and Explanations
(Biology / Life Sciences MCQ: Multiple Choice Questions)
(1). _____ can serve as an alternative to ethidium bromide to stain DNA for detection on gels
a. Mitomycin C
b. Acridine orange
c. SYBR Green I
d. Acriflavine
(2). _____ is a sensitive technique to find out the number of template molecules originally present in a PCR reaction.
a. Hot start PCR
b. Real time PCR
c. AP PCR
d. Reverse Transcriptase PCR
(3). Using __________ it is possible to generate fluorescence in quantitative PCR reactions
a. TaqMan probes
b. Molecular beacons
c. Scorpion probes
d. All of the above
(4). For whole genome amplification, creating very long DNA products ________ method can be employed with great success
a. MDA
b. PEP
c. iPEP
d. DOP
(5). ________ polymerase has the ability to add ~70,000 nucleotides every time it binds to primer and has a very low error rate.
a. Taq polymerase
b. Tli polymerase
c. Pfu polymerase
d. Bacteriophage ϕ 29 polymerase
(6). ________ restriction endonucleases cleaves DNA upto 1000 bp away from the recognition sequence
a. Type I
b. Type II
c. Type III
d. Type IV
(7). _______ is a colourless compound
a. Curcumin
b. Phycocyanin
c. X-gal
d. Carotenes
(8). _____ enzyme can cleave and join DNA molecules
a. Gyrase
b. DNA ligase
c. DNA polymerase
d. Primase
(9). In a covalently closed circular DNA molecule, if linking number (L) is equal to Twist (T) then Writhe is equal to _________
a. Zero
b. Twice (T)
c. ½
d. One
(10). ______ phage is widely used for phage display technique
a. M13
b. Φ 6
c. λ phage
d. T7 phage
(11). ______ RNA polymerase has a single subunit
a. E. coli
b. Bacillus subtilis
c. T7
d. λ phage
(12). Dideoxynucleotides have ________ at their 3’ end
a. CH3
b. H
c. OH
d. NH3
(13). _____ permits sequence analysis in real time
a. Sanger’s classical method of sequencing
b. Maxam and Gilberts method of sequencing
c. Cycle sequencing
d. Pyro-sequencing
(14). Site directed mutagenesis on a DNA molecule can be done using_________.
a. Chemical mutagens
b. Physical mutagens
c. Synthetic olios
d. Random cleavage and ligation
(15). 35s, and mas promoters are used to express genes in _____
a. Mammalian cells
b. Plant cells
c. Bacterial cells
d. Fungal cells
You may also like…
Original Solved Question Papers of DBT BET JRF Exam
| 2016 | 2015 | 2014 | 2013 | 2012 | 2011 | 2010 | 2009 | 2008 |
Answer key with Explanations
1. Ans. (b) Acridine Orange
Ethidium bromide (EtBr) is a DNA intercalating agent commonly used in molecular biology experiments to visualize or locate nucleic acids (DNA or RNA) in agarose gel electrophoresis. It is a fluorescent dye excited by UV light and emits an orange colour. The intensity of the fluorescence of EtBr shows about 20 fold increase when it bound to DNA or RNA and make it an excellent fluorescent tag for the detection of nucleic acids. The absorption maximum of EtBr is 210 nm and 285 nm (UV light). The emission maximum of EtBr is 605 nm (orange) EtBr is a known mutagen to human. EtBr has been used in the treatment of trypanosomiasis in cattle under the trade name homidium.
Acridine orange is a good alternative to EtBr in molecular biology experiments. Acridine orange is more commonly used for the cell cycle stage determination since it can rapidly penetrate into the cells. Acridine binds to DNA by intercalation whereas it binds to RNA by electrostatic attraction.
Excitation and emission maxima of Acridine orange on DNA: Excitation maxima: 502 nm, emission maxima 525 (green) similar to fluorescein.
Excitation and emission of Acridine orange on RNA: Excitation maxima: 460 nm (blue); Emission maxima: 650 nm (red).
Mitomycin C is a chemotherapeutic agent used in the treatment of many types of cancers in human. Mitomycins are group of natural anticancer agents obtained from Streptomyces caespitosus or Streptomyces lavendulae. Three types of mitomycins are so far described and they are named as Mitomycin A, Mitomycin B and Mitomycin C
SYBR Green I (or SG I) is a fluorescent nucleic acid stain used in molecular biology experiments. SYBR Green I is commonly used to quantify DNA in Quantitative Real Time PCRs. It can also be used to visualize DNA in gel electrophoresis and also in flow cytometer.
Acriflavine is a fluorescent dye used in molecular biology to label high molecular weight RNA.
2. Ans. (b). Real Time PCR
Real Time PCR: an advanced PCR technique which allows monitoring of the rate of amplification of template DNA in real time. Exact quantification of the template DNA can be determined with real time PCR methods.
Hot Start PCR: A modified PCR method to reduce the nonspecific amplification of template DNA by the taq polymerase enzyme at lower temperature.
AP PCR or Arbitrarily Primed PCR is commonly known as RAPD (Randomly Amplified Polymorphic DNA). In AP PCR, a short arbitrary primer is used to amplify multiple fragments of DNA from the whole genome isolate of an organism to get a DNA fingerprint. AP PCR fingerprints of different samples can be compared.
Reverse Transcriptase PCR: A modified PCR method to synthesize cDNA from total RNA extracts. It uses Reverse transcriptase enzyme and oligo dT primers. The oligo dT primers will bind to the poly A tail of mRNA which is then extended by the reverse transcriptase enzyme to synthesize the cDNA complementary to the sequence on the mRMA. Thus, with RT PCR total cDNA of an organism can be prepared.
3. Ans. (d). All of the above
4. Ans. (a). MDA
MDA = Multiple Displacement Amplification. MDA is a non PCR based DNA amplification method largely adopted to amplify small amounts of template DNA into large quantities required for the whole genome sequencing projects. Amplification reaction is carried out with a high fidelity polymerase enzyme such as Φ 29 DNA polymeras.
5. Ans. (d). Bacteriophage ϕ 29 polymerase
Taq polymerase:
Obtained from: Thermus aquaticus
Optimum temperature: 75 – 80oC
Replication speed: 1000 bp in 10 seconds
Drawback: lack the 3’5’ exonuclease proof reading acclivity
Error rate: 1 in 9000 nucleotide
Produce ‘A’ overhangs at 3’ ends, thus useful in ‘TA’ cloning with plasmid vector
Tli polymerase
Obtained from Thermococcus litoralis
Pfu polymerase
Obtained from Pyrococcus furiosus
Optimum temperature: 750C
Replication speed: 1000 bp in 1 – 2 minutes
Possess 3’ to 5’ exonuclease proof reading activity
Error rate very less, 1 in 1.3 million base pares
Drawbacks: slow replication rate
Bacteriophage ϕ 29 polymerase
Obtained from Bacteriophage ϕ 29, a phage of Bacillus subtilis
Commonly used in MDA (Multiple Displacement Amplification), a non PCR based DNA amplification process.
Has high affinity for single stranded DNA than double stranded DNA
High processivity, generate large fragments over 10 kb
Thermal cycling process is not required
6. Ans. (a). Type I
Type I: cleave at site away from the recognition site (1000 bp away)
Type II: cleave within the recognition site
Type III: Cleave at sites a short distance from recognition site
Type IV: only target methylated or hydroxymethylated DNA
7. Ans. (c). X-gal
X-gal is a reporter molecule, commonly used to detect the presence of beta galactosidase enzyme in molecular biology experiments. X-gal is colourless compound and which on hydrolysis by the enzyme beta galactosidase yields a blue insoluble compound. The formation of blue colour confirms the presence of active beta galactosidase enzyme.
Curcumin is the active ingredient in the turmeric; it is yellow coloured compound and it is a known anti-cancerous and anti-microbial compound.
Phycocyanin: is the blue pigment of blue green algae (cyanobacteria)
Carotenes: they are yellow or orange coloured plant pigments
8. Ans. (a). Gyrase
DNA gyrase removes the strains of double stranded DNA during its unwinding during DNA replication process.
9. Ans. (a). Zero
10. Ans. (a). M13
11. Ans. (c). T7
12. Ans. (b). H
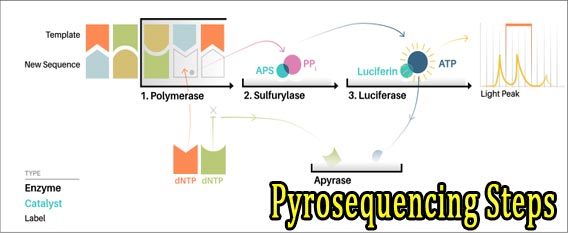
13. Ans. (d). Pyrosequencing
14. Ans. (c). Synthetic Oligos
15. Ans. (b). Plant Cells

Thank you very much for your efforts!