“Genetic variation is important for evolution, but the survival of the individuals
demand genetic stability”
DNA is the genetic information carrier molecule in the cell and thus it is very essential to keep the genetic information intact. Even though DNA holds a prime position, it is one of the highly susceptible molecules in the cells because DNA can be damaged by a number of factors both internal and external in origin. It is very surprising to know that, our cells lose approximately 5000 nucleotides every day due to different damages of the DNA. If these damages are not rectified properly, our cells will be subjected to severe mutations and that will be fatal for the survival of the individual cells and the organism itself. DNA replication process in the cell which ensures the production of exact copy of the genetic information is very accurate due to the high fidelity of DNA polymerase enzyme. However, the process of DNA replication is not 100 percent error free.
What is DNA repair?
DNA polymerase enzyme sometimes accidentally introduces wrong bases which will disrupt the normal Watson-Crick base paring of the DNA. There are also many possibilities of DNA damage during genetic recombination happens during gametogenesis by meiotic cell division. If the damages or errors in the DNA are not corrected in the somatic cells, it may leads to the development of cancer or it results in the loss of function of genes. More than that, if DNA damages occur in the gametes is not rectified, it will be carried over to next generation through progenies. Thus, damage to the genetic materials is a major threat to all organisms. In order to counteract these threats, cells has evolved many methods to overcome and rectify different types DNA damages. All these methods are collectively termed as DNA REPAIR mechanisms. Similar to DNA replication, transcription and translation, the process of DNA repair is also a prime molecular event in the cells which is very essential for the ultimate survival of the cells and also for the survival of the organism.
| You may also like NOTES in... | ||
|---|---|---|
| BOTANY | BIOCHEMISTRY | MOL. BIOLOGY |
| ZOOLOGY | MICROBIOLOGY | BIOSTATISTICS |
| ECOLOGY | IMMUNOLOGY | BIOTECHNOLOGY |
| GENETICS | EMBRYOLOGY | PHYSIOLOGY |
| EVOLUTION | BIOPHYSICS | BIOINFORMATICS |
DNA Repair and Nobel Prize in Chemistry (2015)

The Royal Swedish Academy of Sciences awarded the 2015 Nobel Prize in Chemistry for the discovery and contributions of DNA repair mechanism. The Nobel Prize in Chemistry this year was shared by three scientists namely Thomas Lindhal, Paul Modrich and Aziz Sancar for their “Mechanistic studies of DNA repair”. The detailed mechanism of DNA repair in the cells that we know today is primarily due to their research. Professor Thomas Lindahl demonstrated that the DNA is an unstable molecule which is subjected to damage even under physiological conditions. He also identified a completely new DNA glycosylase enzyme and described their role DNA repair mechanisms.
Professor Paul Modrich transformed the field of mismatch repair to a detailed biochemical understanding first in bacteria and later in eukaryotes. Professor Sancar explained the mechanism of nucleotide excision repair first in bacteria and later in eukaryotic cells. He also explained the molecular mechanisms behind the photoreactivation process, which is type of light dependent DNA repair mechanism. All these contributions helped us to understand the nature of some diseases like cancer and they helped to develop new therapies against many diseases including cancer.
Destructive forces faced by DNA in the cell
There are two categories of destructive forces in the cells that could damage the DNA both structurally and chemically. They are:
(1). Internal or intrinsic factors: they includes:-
Ø Reactive radicals
Ø Metabolic intermediates
Ø Metabolic by-products
Ø Replication errors
Ø Recombination errors
(2). External or extrinsic factors: they includes:-
Ø Radiations (X-rays, UV rays, γ-rays)
Ø Carcinogens/ DNA intercalating agents
As the name suggests, internal factors originate inside the cell itself. Reactive free radicals and metabolic intermediates or metabolic byproducts can severely damage DNA and can induce spontaneous mutations. For example, the oxidative deamination of nucleotides in the cells which results in the conversion of cytosine to uracil is caused by metabolic intermediates. Errors happen during DNA replication and recombination are also considered as the internal factors.
Major external factors that could damage cellular DNA fall under two sub-categories. Among which the different types of radians are most important one. Radiations like UV rays, X-rays and γ-rays can severely disrupt the structure and chemistry of DNA leading to a variety of mutation possibilities. Similarly different carcinogenic chemicals, which we generally called as DNA intercalating agents, can directly react with DNA and can cause different structural and chemical modifications in the DNA.
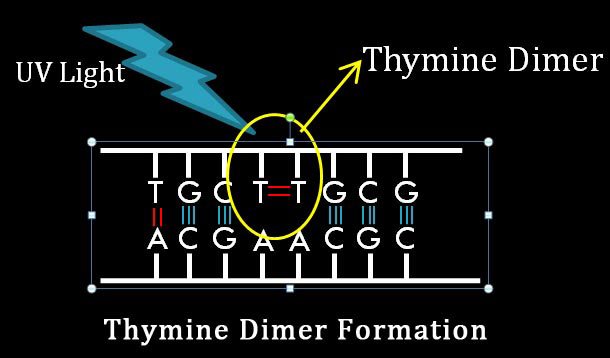
When the normal conformational chemistry of DNA is lost due to any of these internal or external factors, we can say that there is a lesion in the DNA. As in the image, the normal conformational symmetry of DNA is lost due to formation of thymine dimer by UV Light and this created a bulge in one strand. The bulge produced by thymine dimer can be called as DNA lesion.
The possible structural lesions that can happens to DNA:
 (1). Thymine dimer formation:
(1). Thymine dimer formation:
The most common structural lesion in the DNA is the formation of pyrimidine dimer. It is formed by the covalent bond formation between two adjacent pyrimidine residues such as between two thymine or two cytosine or very rarely a thymine and a cytosine. Pyrimidine dimer formation is caused by the ultraviolet irradiation of DNA. Among the three types of pyrimidine dimers, the thymine dimer formation is the most common one.
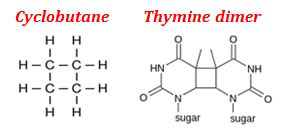
When DNA is struck with UV light, the hydrogen bonds between the two strands breaks and two covalent bonds are formed between the two thymine residues. The two covalent bonds are formed by the breakage of two double bonds present between C5 and C6 of adjacent thymine residues. The Thymine dimer is also called as cyclobutane photodimer or CPD since it structurally resembles cyclobutane nucleus. If thymine dimer in the DNA is left uncorrected, it will cause melanoma, which is a type of skin cancer.
 (2). Spontaneous depurination of DNA
(2). Spontaneous depurination of DNA
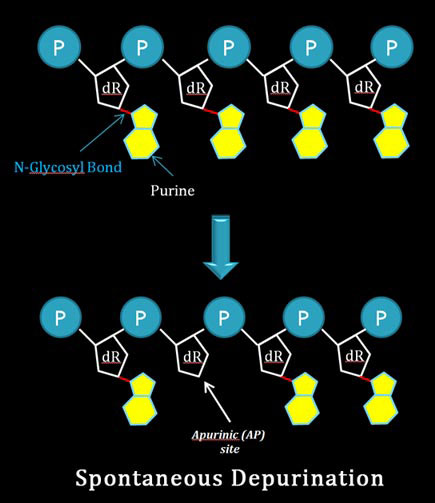
It is due to the spontaneous removal of adenine or guanine residues from the DNA due to the cleavage of N-glycosyl bond which connects the nitrogen base with the deoxyribose sugar. Depurination results in the formation of apurinic site (AP site). Apurinic site structurally disrupt the normal conformation of DNA. If apurinic site is left uncorrected, many types of cancerous growth initiates in the tissue.
(3). Spontaneous deamination of bases in the DNA
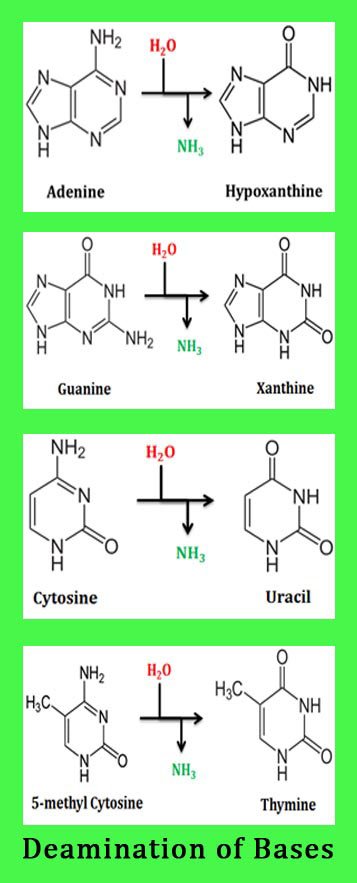 As the name suggests, it the removal of amino groups from the nitrogen bases of DNA. Deamination is usually caused by the oxidative removal of amino group from the nitrogen bases. Deamination produces unnatural bases or change in the base sequence and results in point mutation in the DNA. Unnatural bases are bases other than adenine, guanine, thymine, cytosine or urasil. Hypoxanthine and xanthine are the two unnatural bases incorporated to DNA due to spontaneous deamination.
As the name suggests, it the removal of amino groups from the nitrogen bases of DNA. Deamination is usually caused by the oxidative removal of amino group from the nitrogen bases. Deamination produces unnatural bases or change in the base sequence and results in point mutation in the DNA. Unnatural bases are bases other than adenine, guanine, thymine, cytosine or urasil. Hypoxanthine and xanthine are the two unnatural bases incorporated to DNA due to spontaneous deamination.
Deamination of adenine creates hypoxanthine formation. Deamination of guanine creates another unnatural base called xanthine. Deamination of cytosine creates uracil. Since, uracil is not a DNA base; it will destroy the normal Watson-Crick base pairing of the DNA. Because of the absence of amino group, the deamination of thymine residue is not possible. There is one more type of deamination, in fact it is the most severe and dangerous one. As a part of DNA regulation mechanism, most of the cytosine resides in the DNA will be methylated in its 5th position as 5 methyl cytosine. The deamination of 5-methyl cytosine produces thymine and thus it creates a point mutation.
(4). Errors in DNA replication/Recombination
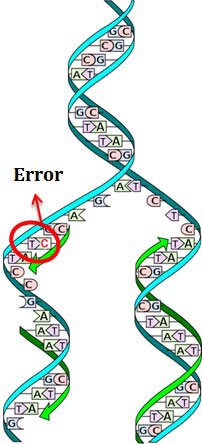
DNA Replication Error
This is due the accidental inclusion of wrong bases in the DNA by DNA polymerase during DNA replication. DNA polymerase also has an exonuclease activity which usually removes the wrong bases and inserts the correct base by a process called ‘proof reading’. Sometimes the proof reading method fails to detect the wrong base and the wrong base persist in the DNA. If these errors are not corrected, in the next replication cycle base change occurs in the DNA and one daughter DNA get mutated.
| You may also like... | ||
|---|---|---|
| NOTES | QUESTION BANK | COMPETITIVE EXAMS. |
| PPTs | UNIVERSITY EXAMS | DIFFERENCE BETWEEN.. |
| MCQs | PLUS ONE BIOLOGY | NEWS & JOBS |
| MOCK TESTS | PLUS TWO BIOLOGY | PRACTICAL |
Different DNA repair mechanisms in the cell:
In this post, we will just mention the names of different DNA repair mechanisms; we have detailed posts with video tutorials for each of these repair mechanisms.
So far there are six different types of DNA repair mechanisms known to science.
(1). Photoreactivation: a light depended DNA repair mechanism which removes thymine dimers
(2). Base Excision Repair (BER): only the damaged base is removed or excised from the DNA strand without removing the normal bases
(3). Nucleotide excision repair (NER): the damaged bases along with a short stretch of healthy stand is removed and refilled with correct bases
(4). Mismatch repair or MMR: As the name suggests it removes the mismatched bases from the DNA and fill with correct bases.
(5). Double strand break repair: double strand break lesions are rectified
(6). Homology directed repair: here a long stretch of DNA repair takes places by consulting with the sequence in the homologous chromosome
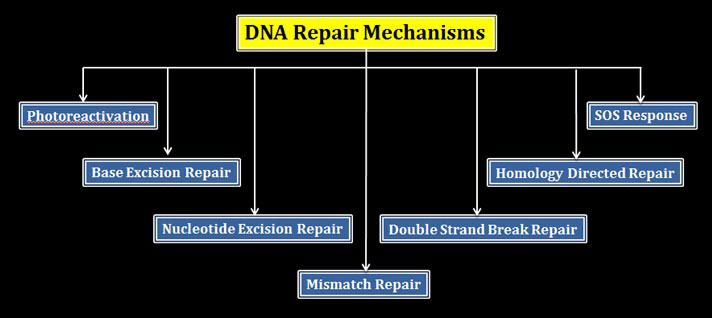
There is another type of DNA repair strategy in the cells called SOS response, is actually not a DNA repair mechanism. SOS response is initiated in the cells after a sever DNA damage. SOS responses trigger many molecular processes in the cells and DNA repair is one among these processes.
The repair mechanisms such as, photoreactivation, base excision repair, nucleotide excision repair and mismatch repair, only the damaged strand of the DNA duplex is repaired and the undamaged strand acts as the template strand. However in double strand break repair and homology directed repair, both the strands of a DNA duplex are repaired.
<<Back to Molecular Biology Notes
