COUNCIL OF SCIENTIFIC & INDUSTRIAL RESEARCH (CSIR) & UGC (Original Question Paper) PART: B (Questions 41 – 55) 41. Which of the following statements about LEAFY (LFY), a regulatory gene in Arabidopsis thaliana, is correct? a. LEAFY (LFY) is involved in floral meristem identity b. LEAFY (LFY) is […]
Continue ReadingMonthly Archives: November 2015
CSIR NET Life Sciences Question Paper: June 2015: with Answer Key and Explanations: Part I
CSIR/JRF/NET: Life Sciences, June 2015 (I) COUNCIL OF SCIENTIFIC & INDUSTRIAL RESEARCH (CSIR) & UGC PART: B (Questions 21 – 40) (21). A 1% (w/v) solution of a sugar polymer is digested by an enzyme (20ug, MW = 200,000). The rate of monomer sugar (MW = 400) liberated was determined […]
Continue ReadingDNA Repair: Part III – Base Excision Repair (BER) Mechanism
Cancer is a disease of the genome. And that’s what happens. You make mistakes in a cell somewhere in your body that causes it to start to grow when it should’ve stopped, and that’s cancer. And those mistakes are mistakes of DNA. Francis Collins What is Base Excision Repair or BER? […]
Continue ReadingPhotoreactivation : Method of DNA Repair for the Recovery of UV Induced DNA Damages by Phytolyase Enzyme and Visible Light
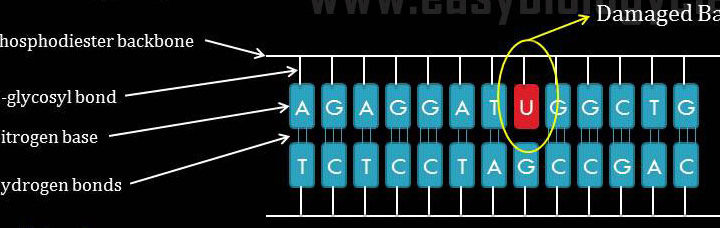
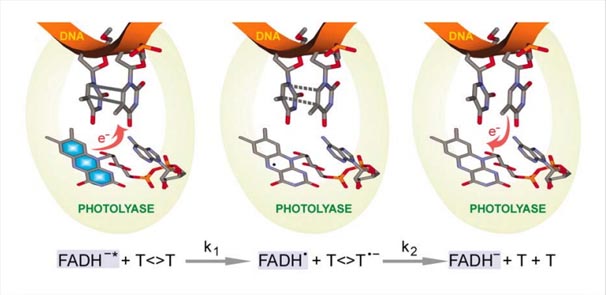
Science is beautiful when it makes simple explanations of phenomena or connections between different observations. Stephen W. Hawking, 2011 What is Photoreactivation? Photoreactivation is a type of DNA repair mechanism present in prokaryotes, archaea and in many eukaryotes. It is the recovery of ultraviolet irradiated damages of DNA by visible […]
Continue ReadingDNA Repair Mechanism – Part I Introduction (DNA Damaging Agents, DNA Damages and Recovery of DNA Damages)
“Genetic variation is important for evolution, but the survival of the individuals demand genetic stability” DNA is the genetic information carrier molecule in the cell and thus it is very essential to keep the genetic information intact. Even though DNA holds a prime position, it is one of the highly […]
Continue Reading