Does the DNA really need to FOLD inside the nucleus?
A diploid human cell contains approximately 6.4 billion base pairs. These 6.4 billion base pairs are distributed in our 23 pairs (2n = 46) of chromosomes. We know that each chromosome contain a single linear segment of DNA.
According to Watson and Crick model, the distance between each base pair in a DNA double helix is 0.34 nm. Thus, the 6.4 billion base pair will constitute a total length of about 2.2 m DNA strand. The total length of DNA of a single human cell is approximately 2.2 meters long (when all 46 DNA strands are joined end to end).
The size of the nucleus in which the chromatin situated is about 10 µm in diameter. Thus, it is evident that the 2.2 m long DNA should fold several times to fit in the nucleus of 10 µm diameter. The exact nature and pattern of folding of DNA strands in the nucleus disclose the organization of genetic material in the cells.
DNA in the nucleus of eukaryotic cells is associated with many proteins. In fact, the compact folding and packing of DNA is facilitated by many proteins which physically interact with DNA. The exact nature of the folding pattern of DNA and its association with proteins were a highly debated topic. Several models are there in the scientific literature regarding the organization of genetic material in the interphase nucleus.
| You may also like NOTES in... | ||
|---|---|---|
| BOTANY | BIOCHEMISTRY | MOL. BIOLOGY |
| ZOOLOGY | MICROBIOLOGY | BIOSTATISTICS |
| ECOLOGY | IMMUNOLOGY | BIOTECHNOLOGY |
| GENETICS | EMBRYOLOGY | PHYSIOLOGY |
| EVOLUTION | BIOPHYSICS | BIOINFORMATICS |
In the previous post, we have discussed the DuPraw Folded Fibre Model of Chromosome. This theory lack solid scientific evidence and hence it is no longer accepted in the scientific community and moreover it is replaced by the more accepted and experimentally proved Nucleosome model of Chromosomes.

Roger D. Kornberg
Nucleosome and Nucleosome Model of Chromosomes
Ø Nucleosome is the lowest level of chromosome organization in eukaryotic cells.
Ø Nucleosome model is a scientific model which explains the organization of DNA and associated proteins in the chromosomes.
Ø Nucleosome model also explains the exact mechanism of the folding of DNA in the nucleus.
Ø It is the most accepted model of chromatin organization.
Ø Nucleosome model of chromosome is proposed by Roger Kornberg (son of Arthur Kornberg) in 1974.
DNA in Eukaryotes are Associated with Histone Proteins:
Ø The chromosomes of eukaryotic organisms composed of DNA and associated proteins.
Ø The DNA and associated proteins are together called as the Chromatin.
Ø The protein components of chromosome (proteins interact with DNA) are grouped into two categories.
(1). Histone proteins
(2). Non-histone proteins
 Ø Histones are basic proteins (positively charged) which contain very large amount of basic amino acids such as lysine and arginine.
Ø Histones are basic proteins (positively charged) which contain very large amount of basic amino acids such as lysine and arginine.
Ø Lysine and arginine are basic amino acids since their side chain (R group) contains additional amino (NH2) groups which can accept protons to form NH3+.
Ø There are five classes of histone proteins in the cells of eukaryotes.
Ø The classification of histones is based on their structural differences, molecular weight and lysine/arginine radio.
Ø The five classes of histones are H1, H2A, H2B, H3 and H4.
Ø The amino acid sequence of histone proteins are highly conserved among organisms.
Ø The reason for this high conserved nature of histone is that, in all organisms the histones are associated with the genetic material, the DNA, which is identical in all the groups.
Ø Nearly all amino acids of the histone protein are in direct contact with the DNA and engaged in interaction with DNA or other histones.
How Scientists understood the Interaction of Histones and DNA in Chromatin?
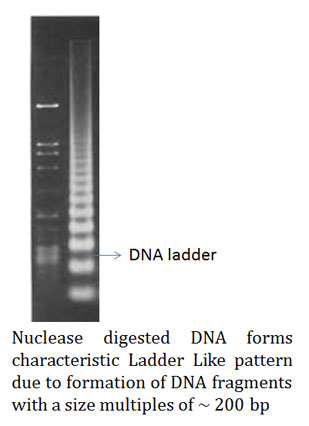
Ø Scientists first treated the chromatin (mixture of DNA and proteins) extracted from a cell with general nuclease enzyme.
Ø After the nuclease digestion, the chromatin sample was run on an agarose gel for its separation.
Ø On the agarose gel, they noticed that the DNA form many fragments of approximately 200 bps or multiples of 200 base pairs.
Ø In contrast to this, when necked DNA (DNA not associated with proteins) was treated with the same nuclease enzyme the fragments generated ware randomly sized.
Ø This suggests the protection of DNA from nuclease enzyme at certain places along its entire length.
Ø Later it was hypothesized that the proteins interacting with the DNA in the chromatin are providing this type of protection.
Ø Based on these findings, Roger Kornberg proposed a model of organization of chromatin called Nucleosome Model.
Nucleosome Model of Chromosome Organization (Interaction of Histones and DNA in Chromatin)
Ø Roger Kornberg proposed that DNA and histones were organized into repeated units called nucleosome.
Ø Nucleosome model is the most accepted model of chromatin.
Ø Nucleosomes are the fundamental repeating units of chromatin.
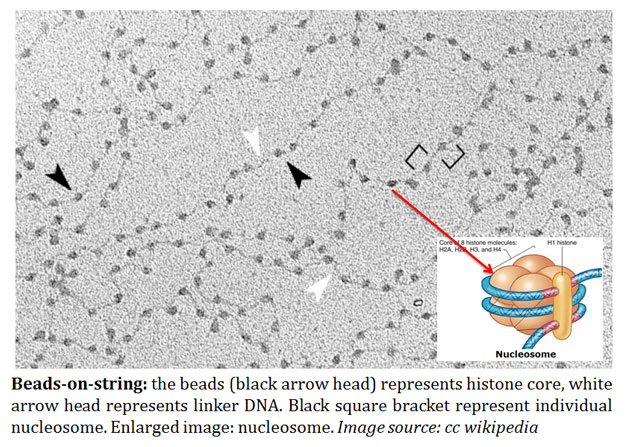
image source: cc wikipedia
Ø Nucleosome represents the ‘beads’ as proposed in the ‘beads on string’ organization of chromatin.
Ø Each nucleosome contains a nucleosome core particle.
Ø The nucleosome core composed of a disc shaped structure of eight histone proteins.
Ø The nucleosome core composed of two molecules of each of the four histones H2A, H2B, H3 and H4 and his structure is called the histone octamer.
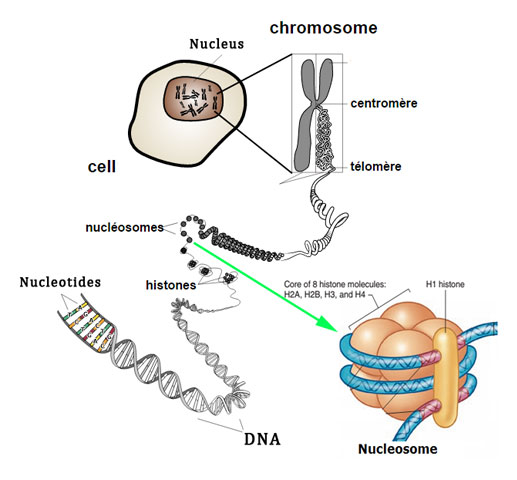
image source: cc wikipedia
Ø The DNA helix is wrapped as super helical left handed turn around this histone octamer core.
Ø Each histone core is encircled by 1.8 turns of DNA.
Ø This 1.8 turn of DNA represents about 146 base pairs.
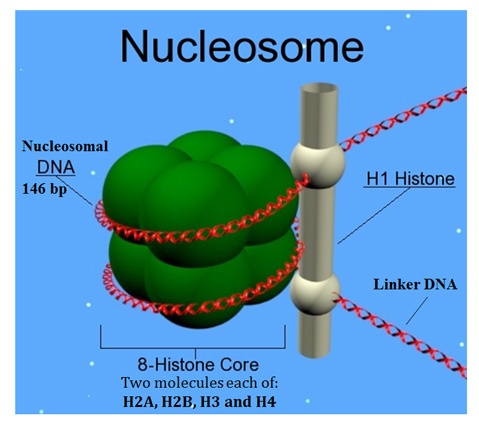
(image source: cc wikipedia)
Ø Each nucleosome is about 10 nm in diameter.
Ø The H1 histone stays outside the histone octamer.
Ø Adjacent nucleosomes are connected by a short stretch of DNA called linker DNA.
Ø Linker DNA is about 10 to 80 bp in length.
Ø H1 histones bind to the liner DNA.
Ø H1 histone binds near the site where DNA enters and exits the nucleosome.
Ø The interaction of histones and DNA in nucleosome is stabilized by several types of non-covalent bonds.
Ø Among these bonds, the ionic bonds formed between the negatively charged phosphate groups in the DNA with the positively charged amino groups of histones were very important.
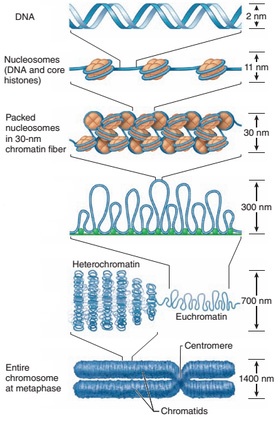
Formation of Metaphase Chromosome
Ø DNA in the chromatin attain a packing ratio of about 7:1 (seven fold packing) by the formation of nucleosomes.
Ø Nucleosome units organized into more compact structure of 30 nm in diameter called 30 nm fibers (proposed by Rachel Horowitz & Christopher Woodcock in 1990).
Ø The H1 histone plays very important role in the formation of the 30-nm fiber.
Ø The formation of 30 nm fiber shortens genetic material (DNA) another seven-fold.
Ø The linker DNA regions in 30-nm structure are variably bent and twisted to attain the folding pattern.
Ø This 30 nm fibres are further folded to form the metaphase chromosome during cell division
<<Back to Cell & Molecular Biology Notes
