Cancer is a disease of the genome. And that’s what happens. You make mistakes in a cell somewhere in your body that causes it to start to grow when it should’ve stopped, and that’s cancer. And those mistakes are mistakes of DNA.
Francis Collins
What is Base Excision Repair or BER?
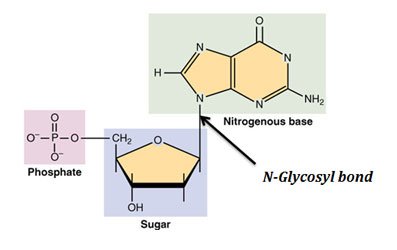
As the name suggests, it is a type of DNA repair mechanism present in both prokaryotes and eukaryotes. In this DNA repair method, the damaged or unnatural base in the DNA double helix is removed by cleaving the N-glycosyl bond without disrupting the phosphodiester bond. N-glycosyl bond is the covalent bond which connects the nitrogen base with the deoxy-ribose sugar of the DNA. The importance of this bond is that, if an enzyme can cleave this particular bond, as it happens during base excision repair, it can selectively excise the nitrogen base from the DNA without altering the phosphodiester backbone.
The main difference of Base Excision Repair from other repair mechanisms is that here only the damaged base is excised from the DNA strand, the phosphodiester back bone is not disturbed for the removal or the damaged bases. But in other DNA repair mechanisms such as mismatch repair or nucleotide excision repair, the damaged nucleotide (nucleotide = nitrogen base + sugar + phosphate group) as such is removed first and refilled by with correct nucleotides. Dear students, please remember, the cleavage of phosphate back bone is also occurs here but it happens in the second stage, not as the part of the removal of nitrogen base.
| You may also like NOTES in... | ||
|---|---|---|
| BOTANY | BIOCHEMISTRY | MOL. BIOLOGY |
| ZOOLOGY | MICROBIOLOGY | BIOSTATISTICS |
| ECOLOGY | IMMUNOLOGY | BIOTECHNOLOGY |
| GENETICS | EMBRYOLOGY | PHYSIOLOGY |
| EVOLUTION | BIOPHYSICS | BIOINFORMATICS |
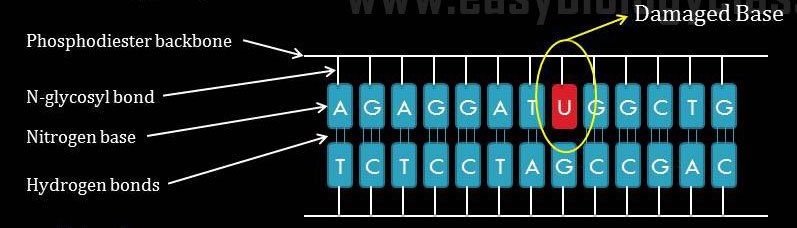
Discovery of Base Excision Repair method:

Tomas Lindahl
The base excision repair mechanism for the first time was described by Professor Tomas Lindahl for which he shared the 2015 – Nobel Prize in chemistry along whit Paul Modrich and Aziz Sancar. Professor Lindahl found that, uracil, an RNA base, is frequently found in the DNA.
Based on this observation he hypothesized that, there must be a mechanism in the cell to handle this problem. As a support of this hypothesis, he discovered a completely new enzyme called uracil DNA glycosylase or UNG and explained their role in the removal of uracil from the DNA. Few years later, with series of experiments, Professor Lindahl explained the biochemistry and molecular mechanism of Base Excision Repair first in bacteria and later in eukaryotes.
Different types of DNA lesions which are rectified by Base Excision Repair method:
Base excision repair mechanism can rectify many single base changes in the DNA. Among these single base changes, the de-aminated bases and alkylated bases are the most common ones. Spontaneous deamination of bases creates unnatural bases in the DNA, It also cause change in base sequence of the DNA. For example, the deamination of adenine creates hypoxanthine, an unnatural base. Another unnatural base called xanthine is created by the deamination of guanine.
Similarly, deamination of cytosine causes uracil formation. Formation of uracil in the DNA is one of the most severe types of deamination in the cellular DNA. Alkylations of bases are another major problem. Alkylation results in the formation chemically different bases which will disrupt the normal Watson-Crick base pairing of the DNA. Examples of alkylation products are 3-methyladenine and 7-methylguanosine. Another major single base abnormality in the DNA is caused by the oxidation of bases. Formation of 8-oxoguanine is the most common product of oxidation of bases in the DNA. All these different types of single base structural lesions can be effectively rectified by base excision repair mechanism.
Enzymes involved in Base Excision Repair Method:
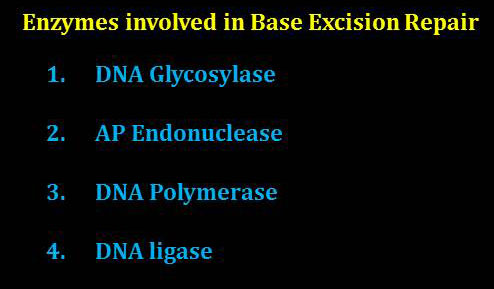 Mainly four different enzymes are involved in the base excision repair mechanism of DNA. They are:
Mainly four different enzymes are involved in the base excision repair mechanism of DNA. They are:
(1). DNA gycosylase: The enzyme which cleaves the N-glycosyl bond between the damaged base and the deoxy-ribose sugar. There are specific DNA glycosylases for each type of damaged bases. Among which the Uracil DNA glycosylase or UNG is the most abundant one and it also the first isolated DNA glycosylase enzyme by Professor Thomas Lindhal. Uracil DNA glycosylase or UNG removes the Uracil residue form the DNA. UNG is very specific for the uracil residue present in the double stranded DNA. It cannot act on the uracil molecule of RNA or single stranded DNA.
(2). AP endonuclease: It is a special type of endonuclease enzyme. ‘AP’ stands for apurinic or apyrimidinic, which means a portion of DNA without any purine or pyrimidine. As we mentioned earlier, the removal of damaged base with DNA glycosylase enzyme by the cleavage of N-glycosyl bond creates an AP site in the DNA. The AP endonuclease recognizes the AP site and specifically cut the phosphodiester back bone at the 5’ portion.
(3). DNA polymerase enzyme: This enzyme fills the gap of DNA created by the removal of damaged bases in the DNA by the action of DNA glycosylase and AP endonuclease enzyme. In prokaryotes, the gap filling as a part of base excision repair is done by DNA polymerase I where as in eukaryotes, it is done by DNA polymerase β.
(4). DNA ligase: Ligase seals the nick in the DNA strand after the polymerase enzyme fills the gaps.
The simultaneous and orderly action of these four enzymes is very essential for the proper orchestration of base excision repair mechanism in the cell.
Molecular Mechanism of Base excision repair:
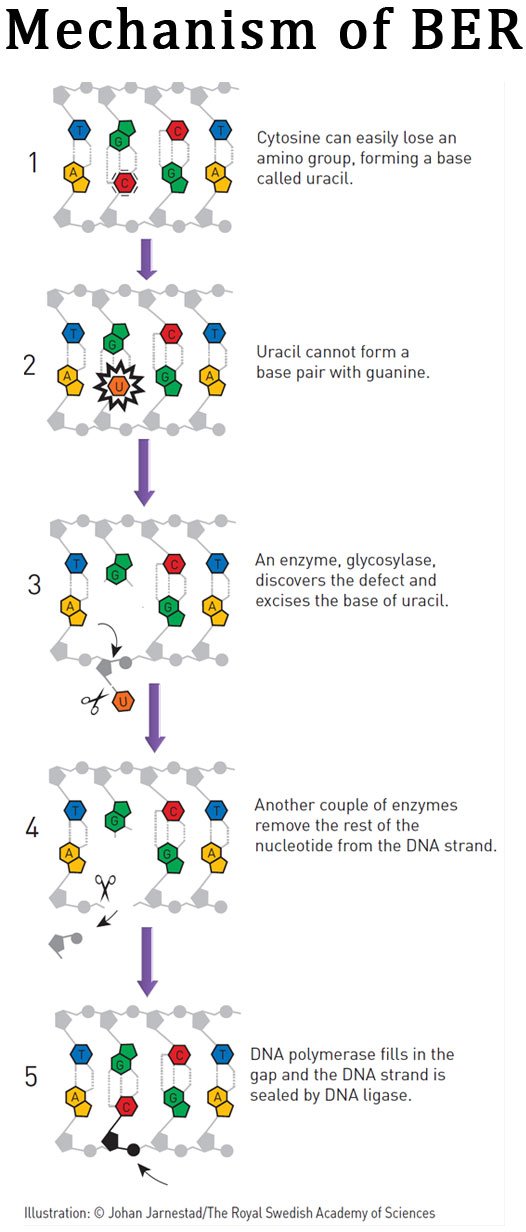 The most cited base excision repair mechanism in the cell is the one which removes the uracil residue from the DNA by the action of the enzyme uracil DNA glycosylase or UNG. Uracil in the DNA is created by the spontaneous deamination of cytosine. Since uracil is not the DNA base, once uracil is formed in the DNA it will destroy the normal Watson-Crick base pairing. Thus, it should be removed from the DNA, otherwise it may leads to mutations or strand breaks in the DNA. To tackle this problem, cell has an enzyme called uracil DNA glcosylase or UNG which removes the uracil residue from the DNA.
The most cited base excision repair mechanism in the cell is the one which removes the uracil residue from the DNA by the action of the enzyme uracil DNA glycosylase or UNG. Uracil in the DNA is created by the spontaneous deamination of cytosine. Since uracil is not the DNA base, once uracil is formed in the DNA it will destroy the normal Watson-Crick base pairing. Thus, it should be removed from the DNA, otherwise it may leads to mutations or strand breaks in the DNA. To tackle this problem, cell has an enzyme called uracil DNA glcosylase or UNG which removes the uracil residue from the DNA.
The Uracil glycosylase enzyme scans the DNA and if it found a uracil reside in the DNA, it will cleave the N-glycosyl bond between the uracil and its deoxy-ribose sugar. As a result of this, the uracil residue is cleaved out from the DNA without disturbing the phosphodiester back bone. This type of removal of uracil creates a region in the DNA without a pyrimidine and as we mentioned earlier, this regions is called the AP site.
The AP site is recognized by a special endonuclease enzyme called AP endonuclease. AP endonuclease cleaves at the 5’ region on the phosphate back bone of the AP site and the sugar phosphate is cleaved off from the DNA. Then the repair DNA polymerase recognizes this DNA strand and fills with correct nucleotide, (here a cytosine residue). In prokaryotes the DNA polymerase I is the enzyme involved in base excision repair mechanism, whereas in eukaryotes, the gap filling is done by DNA polymerase beta. Then finally a special type of DNA ligase enzyme called DNA ligase III which seals the nick created by the addition of new nucleotide by the DNA polymerase enzyme. Thus, with this repair mechanism, the original sequence of the DNA is regained.
Similar to the base Excision repair mechanism for uracil, there are also methods for other types of damaged bases or un-natural bases incorporated in DNA such as for the removal of oxides of bases, alkylated bases, or bases produced by spontaneous deamination such as xanthine and hypoxanthine.

Significance / Importance of Base Excision Repair:
Base excision repair is important to remove damaged bases from the DNA. The damaged bases if persist in the DNA, it can create mutation my miss pairing or leads to DNA breaks during DNA replication. Thus existence of base excision repair mechanism is very essential for the ultimate survival of the individuals. It is fond that mutation in the genes of base excision repair machinery in human can leads to the development of a variety of cancer including colon cancer.
Key points of base excision repair mechanism:
Ø Base excision repair or BER is a DNA repair method present in both prokaryotes and eukaryotes.
Ø BER specifically removes the damaged based from the DNA by cleaving the N-glycosyl bond.
Ø The BER machinery and its molecular mechanism were first described by Professor Tomas Lindahl.
| You may also like... | ||
|---|---|---|
| NOTES | QUESTION BANK | COMPETITIVE EXAMS. |
| PPTs | UNIVERSITY EXAMS | DIFFERENCE BETWEEN.. |
| MCQs | PLUS ONE BIOLOGY | NEWS & JOBS |
| MOCK TESTS | PLUS TWO BIOLOGY | PRACTICAL |
Ø The BER mechanism includes four enzymes, they are
Ø DNA glycosylase which cleaves the N-glycosyl bond between the damaged base and the sugar to remove the damaged base from the DNA. There are specific DNA glycosylases for each of the base damages. Uracil DNA glycosylase is the most abundant one.
Ø AP endonuclease which cut the phosphate backbone at the AP site.
Ø DNA polymerase which fills the gap in the DNA with correct base.
Ø DNA ligase enzyme which seals the nick.
Ø BER machinery is very essential to maintain the integrity of genomic DNA in the cell.
Ø Mutations in the any of the genes of BER machinery may create mutations.
<<Back to Molecular Biology Notes
